
Recombinase Polymerase Amplification Information & Publications
At Intact Genomics, we lead the development of both PCR and RPA enzyme products to give you the highest quality enzymes available. Our experienced Research and Development team drives our dedication to making reliable molecular biology products for our customers. Although PCR has been around for decades, RPA emerges as a developing technology that has the potential to change how molecular biology is performed across many disciplines.
Recombinase Polymerase Amplification (RPA) is an enzyme- based alternative to Polymerase Chain Reaction (PCR). Like PCR, the primary objective of RPA is to make abundant copies of a targeted DNA sequence. Both PCR and RPA use a set of primers that target a DNA sequence of interest and a polymerase to make copies of that DNA sequence (Figure 1). RPA achieves maximum amplification faster than PCR and the reaction takes place at a single temperature, usually 37 °C. Since the reaction is chemistry based, the technology can be adapted to portable kits or test tubes without the need for the lab-based thermocyclers that are required with PCR. Advances in RPA research and production are quickly approaching and surpassing most standards set by PCR.
In the 1960s, Walter Harm found that UV-sensitive (i.e. “uvs”) bacteriophage T4 mutants had altered enzymes that affected their ability to repair host double-stranded DNA. Later groups isolated uvsX and other genes that could repair broken double-stranded DNA using single stranded DNA. This same mechanism—a protein combining double-stranded DNA with single-stranded DNA—is the key to RPA. UvsX together with DNA binding proteins uvsY, and gp32 (“gene protein”) work together to recruit single-stranded DNA primers and open DNA sites for a polymerase to make copies of your DNA.
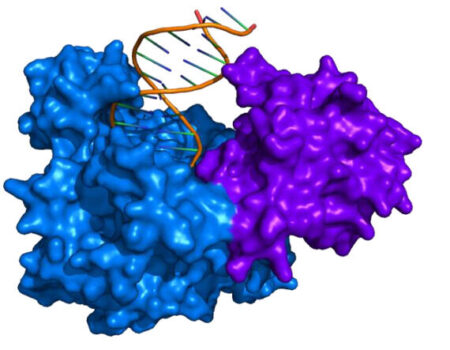
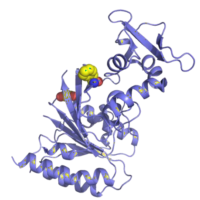
In addition to a polymerase (typically Bsu or Sau), RPA enzymes include the DNA recombinase (UvsX) and DNA binding proteins (UvsY and gp32). A structural representation of UvsX is shown in Figure 2. Like RecA, UvsX can simultaneously bind both double-stranded DNA and a single stranded DNA. The UvsX-dsDNA-primer complex can have several UvsX molecules that work together with UvsY and gp32 to direct polymerase activity. This is all you need to amplify DNA!
We are striving to provide you with even better tools for your research. Keep an eye on our products as we improve upon existing technologies.
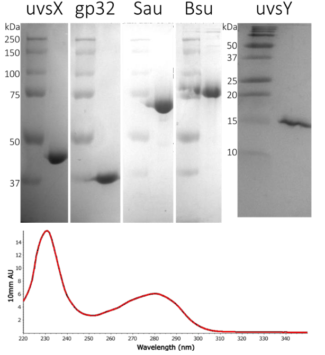
RPA requires multiple high-quality enzymes, whereas PCR only needs one pure polymerase. Our enzymes are produced biologically in E. coli, which easily produce large quantities of enzymes. All biological sources of proteins produce many other proteins that the bacteria need to survive. Separating the enzymes that we need from all the other bacterial proteins is essential for RPA, especially since bacteria produce DNA and RNA nucleases that can inhibit the amount of amplification. At Intact Genomics, we do that work so you don’t have to. We purify, prepare and test the enzymes for you to get to work right away (Figure 3). Intact Genomics provides you with the enzymes at the concentrations where the activities are optimized for you to get started with RPA. We also make sure that our enzymes are nuclease free so that amplification and detection are robust measurements.
Our RPA research is ongoing, therefore keep following us as we innovate new products in several areas of molecular biology to help you perform your research faster and easier.
Intact Genomics RPA Related Products
Intact Genomics offers a variety of RPA optimization products. Each product is put through rigorous QC (for activity, concentration and purity) and can be customized. Bulk order discount is available. Please contact us at sales@intactgenomics.com.
| Product Links | Package Size | Features |
| IG® RPA Master Mix | 25 rxns, 100 rxns, 500 rxns | IG RPA Master Mix delivers powerful isothermal amplification for your molecular diagnostics and research. Easy-to-use, fast, reliable, and more affordable. |
| ig® Recombinase Polymerase Amplification (RPA) Kit v2 | 25 rxns, 100 rxns, 500 rxns | The new improved IG® Recombinase Polymerase Amplification Version 2 Kit (IG® RPA v2) helps researchers who seek the advantages of isothermal DNA amplification to achieve faster, more discernible results. |
| FastAmp® qRPA SYBR kit | 25 rxns, 100 rxns, 500 rxns | FastAmp® qRPA SYBR Kit is a powerful tool to screen primer conditions for the lowest possible Non-template control (NTC) background. |
| FastAmp® Plant Direct RPA kit | 25 rxns, 100 rxns, 500 rxns | FastAmp® Plant Direct RPA kit offers a fast, robust method for amplifying DNA fragments from plant tissues at a single and constant temperature (37°C) using RPA technology. |
| ig® Recombinase Polymerase Amplification (RPA) Kit | 25 rxns, 100 rxns, 500 rxns | This introductory kit gives the user a faster, more streamlined Recombinase Polymerase Amplification experience by providing the necessary reagents and enzymes to amplify user-provided DNA template. |
| T4 UvsX DNA Recombinase | 100 μg, 500 μg, & 1000 µg | Important for the error-free repair of DNA double-strand breaks and for replication fork restart |
| T4 UvsY Protein | 100 μg, 500 μg, & 1000 µg | T4 UvsY is the phage T4 recombination mediator protein. It stimulate sthe filament nucleation event |
| T4 gp32 Protein | 100 µg, 200 µg, 500 μg | Single-stranded DNA binding protein required for T4 DNA replication, recombination, and repair |
| Bsu DNA Polymerase Large Fragment | 1,000 units | Polymerase useful for extending primers in RPA |
| Sau DNA Polymerase Large Fragment | 1,000 units | Polymerase useful for extending primers in RPA |
| Glycerol Free T4 UvsX DNA Recombinase | 100 μg, 500 μg, & 1000 µg | Important for the error-free repair of DNA double-strand breaks and for replication fork restart |
| Glycerol Free T4 UvsY Protein | 100 μg, 500 μg, & 1000 µg | T4 UvsY is the phage T4 recombination mediator protein. It stimulate sthe filament nucleation event |
| Glycerol Free T4 gp32 Protein | 500 μg | Single-stranded DNA binding protein required for T4 DNA replication, recombination, and repair |
| Glycerol Free Bsu DNA Polymerase Large Fragment | 1,000 units | Polymerase useful for extending primers in RPA |
| Exonuclease III | 10000 units, 25000 units | Exonuclease III (Exo III) is an exodeoxyribonuclease that digests one strand of duplex DNA from a blunt end, 5′ overhang or nick |
| Endonuclease IV(Nfo) | 2000 units, 5000 units | Endonuclease IV (Nfo) from Escherichia coli is a 32-kD metalloprotein that aids in the repair of damaged DNA. The enzyme functions both as an apurinic/apyrimidinic nuclease and as a 3′-terminal di-esterase |
| FastAmp® Viral and Cell Lysis Solution (1x, 5x) | 25 reactions, 100 reactions | FastAmp® Viral and Cell Lysis Solution streamlines DNA and RNA collection and storage processes. The lysed DNA or RNA can be directly used in RPA, RT-isothermal DNA amplification, PCR and RT-qPCR, , without the need of an extraction step. |
Select Publications for Further Study
If you are new to RPA and want to know more, here are a few references to help you get started1–34. We want to help you improve your productivity.
- Ozay, B. & McCalla, S. A review of reaction enhancement strategies for isothermal nucleic acid amplification reactions. Sens. Actuators Rep. 3, 100033 (2021).
- Harm, W. On the control of UV-sensitivity of phage T4 by the gene x. Mutat. Res. Mol. Mech. Mutagen. 1, 344–354 (1964).
- Mukama, O. et al. An ultrasensitive and specific point-of-care CRISPR/Cas12 based lateral flow biosensor for the rapid detection of nucleic acids. Biosens. Bioelectron. 159, 112143 (2020).
- Liu, J. & Morrical, S. Assembly and dynamics of the bacteriophage T4 homologous recombination machinery. Virol. J. 7, 357 (2010).
- Carles-Kinch, K., George, J. & Kreuzer, K. Bacteriophage T4 UvsW protein is a helicase involved in recombination, repair and the regulation of DNA replication origins. EMBO J. 16, 4142–4151 (1997).
- Gajewski, S. et al. Crystal Structure of the Phage T4 Recombinase UvsX and Its Functional Interaction with the T4 SF2 Helicase UvsW. J. Mol. Biol. 405, 65–76 (2010).
- Piepenburg, O., Williams, C., Stemple, D. & Armes, N. DNA Detection Using Recombination Proteins. PLOS Biol. 4, e204 (2006).
- Derbyshire, V. et al. Genetic and Crystallographic Studies of the 3’,5’-Exonucleolytic Site of DNA Polymerase I. Science 240, 199–201 (1988).
- Perumal, S., Nelson, S. & Benkovic, S. Interaction of T4 UvsW helicase and single-stranded DNA binding protein gp32 through its carboxy terminal acidic tail. J. Mol. Biol. 425, 2823–2839 (2013).
- Van Ness, J., Van Ness, L. & Galas, D. Isothermal reactions for the amplification of oligonucleotides. PNAS 100, 4504–4509 (2003).
- Liu, J., Berger, C. & Morrical, S. Kinetics of Presynaptic Filament Assembly in the Presence of Single-Stranded DNA Binding Protein and Recombination Mediator Protein. Biochemistry 52, 7878–7889 (2013).
- Steinegger, M. & Soding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
- Li, J., Macdonald, J. & von Stetten, F. Review: a comprehensive summary of a decade development of the recombinase polymerase amplification. Analyst 144, 31–67 (2019).
- Sweezy, M. & Morrical, S. Single-stranded DNA Binding Properties of the uvsY Recombination Protein of Bacteriophage T4. J. Mol. Biol. 266, 927–9358 (1997).
- Story, R., Bishop, D., Kleckner, N. & Steitz, T. Structural Relationship of Bacterial RecA Proteins to Recombination Proteins from Bacteriophage T4 and Yeast. Science 259, 1892–1896 (1993).
- Yonesaki, T. & Minagawa, T. T4 phage gene uvsX product catalyzes homologous DNA pairing. EMBO J. 4, 3321–3327 (1985).
- Sickmier, E. A., Kreuzer, K. & White, S. The Crystal Structure of the UvsW Helicase from Bacteriophage T4. Structure 12, 583–592 (2004).
- Aliotta, J. et al. Thermostable Bst DNA polymerase I lacks a 3’+ 5’ proofreading exonuclease activity. Genet. Anal. Biomol. Eng. 12, 185–195 (1996).
- Bleuit, J. S., Ma, Y., Munro, J. & Morrical, S. W. Mutations in a Conserved Motif Inhibit Single-stranded DNA Binding and Recombination Mediator Activities of Bacteriophage T4 UvsY Protein. J. Biol. Chem. 279, 6077–6086 (2004).
- Chen, W. et al. Combining simplified DNA extraction technology and recombinase polymerase amplification assay for rapid and equipment-free detection of citrus pathogen Phytophthora parasitica. J. Integr. Agric. 20, 2696–2705 (2021).
- Formosa, T. & Alberts, B. M. Purification and characterization of the T4 bacteriophage uvsX protein. J. Biol. Chem. 261, 6107–6118 (1986).
- Ishmael, F. T., Alley, S. C. & Benkovic, S. J. Identification and Mapping of Protein-Protein Interactions between gp32 and gp59 by Cross-linking. J. Biol. Chem. 276, 25236–25242 (2001).
- Jauset-Rubio, M. et al. Ultrasensitive, rapid and inexpensive detection of DNA using paper based lateral flow assay. Sci. Rep. 6, 37732 (2016).
- Kerr, I. D. et al. Crystallographic and NMR Analyses of UvsW and UvsW.1 from Bacteriophage T4. J. Biol. Chem. 282, 34392–34400 (2007).
- Lillis, L. et al. Factors influencing Recombinase polymerase amplification (RPA) assay outcomes at point of care. Mol. Cell. Probes 30, 74–78 (2016).
- Lovett, S. T. The DNA Exonucleases of Escherichia coli. EcoSal Plus 4, ecosalplus.4.4.7 (2011).
- Mekuria, T. A., Zhang, S. & Eastwell, K. C. Rapid and sensitive detection of Little cherry virus 2 using isothermal reverse transcription-recombinase polymerase amplification. J. Virol. Methods 205, 24–30 (2014).
- Munawar, M. A., Martin, F., Toljamo, A., Kokko, H. & Oksanen, E. RPA-PCR couple: an approach to expedite plant diagnostics and overcome PCR inhibitors. BioTechniques 69, 270–280 (2020).
- Nelson, S. W. & Benkovic, S. J. The T4 Phage UvsW Protein Contains Both DNA Unwinding and Strand Annealing Activities. J. Biol. Chem. 282, 407–416 (2007).
- Nuntawong, P., Putalun, W., Tanaka, H., Morimoto, S. & Sakamoto, S. Lateral flow immunoassay for small-molecules detection in phytoproducts: a review. J. Nat. Med. 76, 521–545 (2022).
- Salazar, A., Ochoa-Corona, F. M., Talley, J. L. & Noden, B. H. Recombinase polymerase amplification (RPA) with lateral flow detection for three Anaplasma species of importance to livestock health. Sci. Rep. 11, 15962 (2021).
- Webb, M. R., Plank, J. L., Long, D. T., Hsieh, T. & Kreuzer, K. N. The Phage T4 Protein UvsW Drives Holliday Junction Branch Migration. J. Biol. Chem. 282, 34401–34411 (2007).
- Wu, H. et al. A Recombinase Polymerase Amplification and Lateral Flow Strip Combined Method That Detects Salmonella enterica Serotype Typhimurium With No Worry of Primer-Dependent Artifacts. Front. Microbiol. 11, 1015 (2020).
- Yonesaki, T., Miyazakie, J. & Minagawa, T. Genetic Studies on Non-Lethal Recombination Gene Mutants of Bacteriophage T4. Mem. Fac. Sci. Kyoto Univ. Ser. Biol. 9, 87–96 (1984).
